COVID-19 dominated the news cycle of 2020. SARS-CoV-2 variants took over news feeds in 2021. For the past year, researchers around the globe have been coming together to openly share data and resources, fighting the pandemic from a unified front. The global investigation effort requires constant update on the status of different emerging lineages and mutations, and this is a huge challenge. In the US alone, states with varied rates of sequencing and reporting, the lack of a national variant surveillance system, and an overwhelming amount of constantly changing information pouring in from numerous sources makes it difficult to know what is going on at any given moment.
The SARS-CoV-2 Mutation Reports built into the Outbreak.info dashboard help solve this problem. Using the Lineage|Mutation Tracker, the Location Tracker, and the Lineage Comparison tool, we can get an impressive overview of the variants, including Variant of Concern B.1.1.7.
The B.1.1.7 lineage, initially labeled Variant of Concern 202012/01, or colloquially the “UK variant,” was first detected in the United Kingdom in samples dated September 20, 2020. In 75% of sequenced samples, this lineage contains 9 mutations in the virus’s S-gene (the spike protein that helps the virus to dock on human cells), including Mutations of Interest N501Y and P681H. N501Y frequently shows up in the B.1.351 (“South Africa variant”) and P.1("Brazil variant") lineages. This mutation in the spike protein has shown to increase the transmissibility of the virus. Two deletion mutations in the B.1.1.7 lineage at the 69th and 70th amino acids have also shown to make the virus more transmissible.
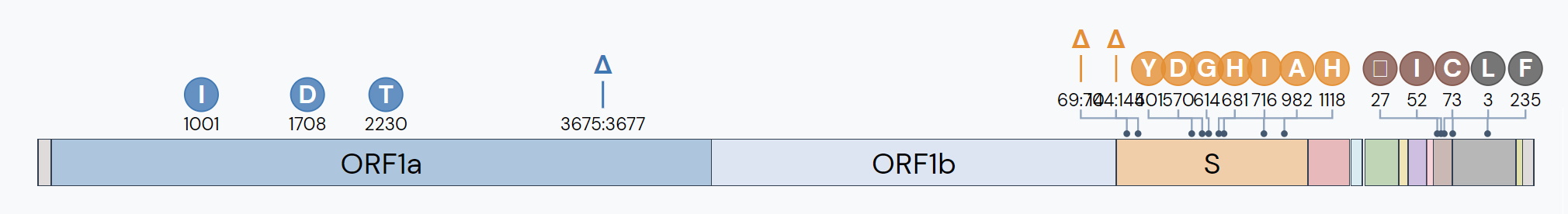
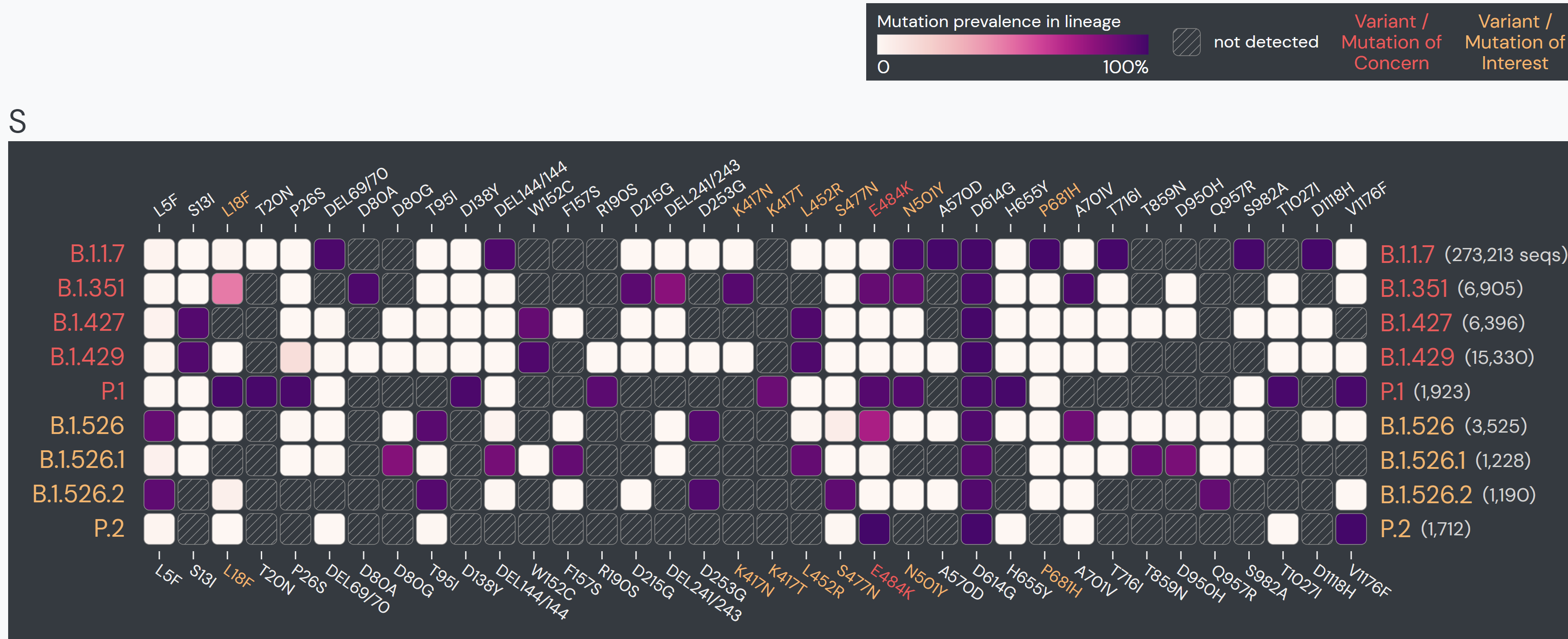
In December 2020, the first sequence of B.1.1.7 lineage with the E484Kmutation was detected. This caused some alarm, as E484K is a Mutation of Concern that has shown to reduce antibody binding. However, to date, this combination makes up less than half a percent of B.1.1.7 sequences.
Before discussing prevalence, it must be noted B.1.1.7 prevalence is over-estimated due to sequencing bias. Most B.1.1.7 genomes identified in the United States have been identified by S-gene target failures (SGTF) in community-based diagnostic PCR testing. SGTF indicates a deletion mutation that is one of several mutations able to distinguish the B.1.1.7 from other strains. The lag time between the sample being acquired, sequenced, and added to the database also means the data for recent dates usually contains very few samples, limiting its reliability. Click here to read more about sequencing bias and limitations.
The lineage emerged in the UK in late September 2020. By November, about 25% of sequenced samples in England were in the B.1.1.7 lineage. By mid-December, this rose to 60%.
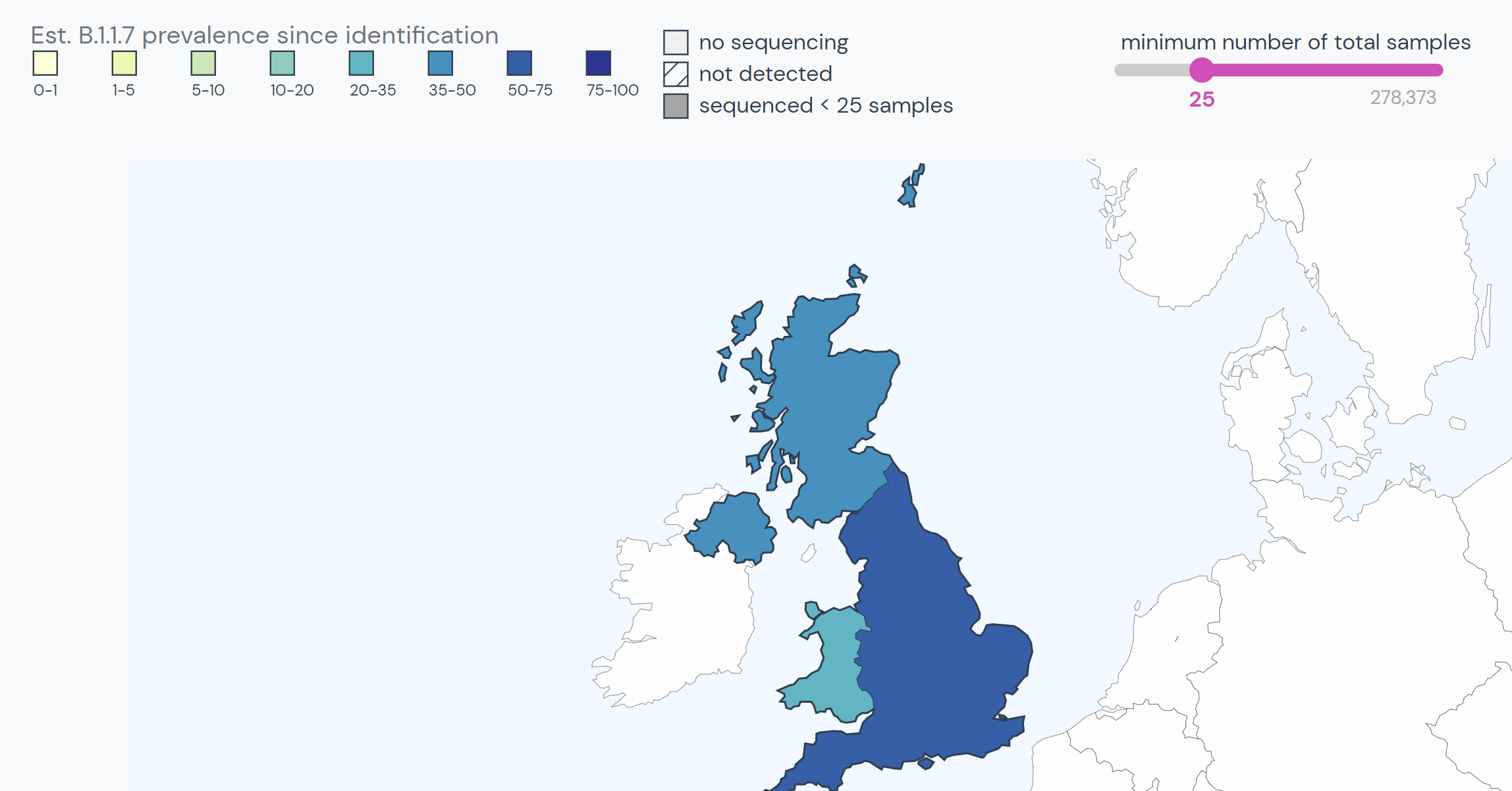
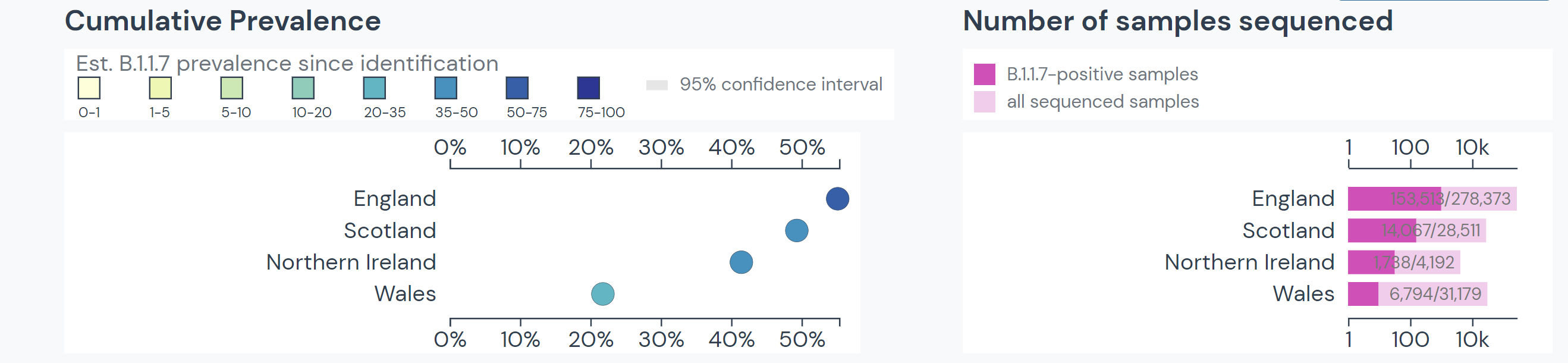
Since its discovery in the UK, the lineage has spread to at least 108 countries and all US states. It is the dominant SARS-CoV-2 variant in over 20 countries.
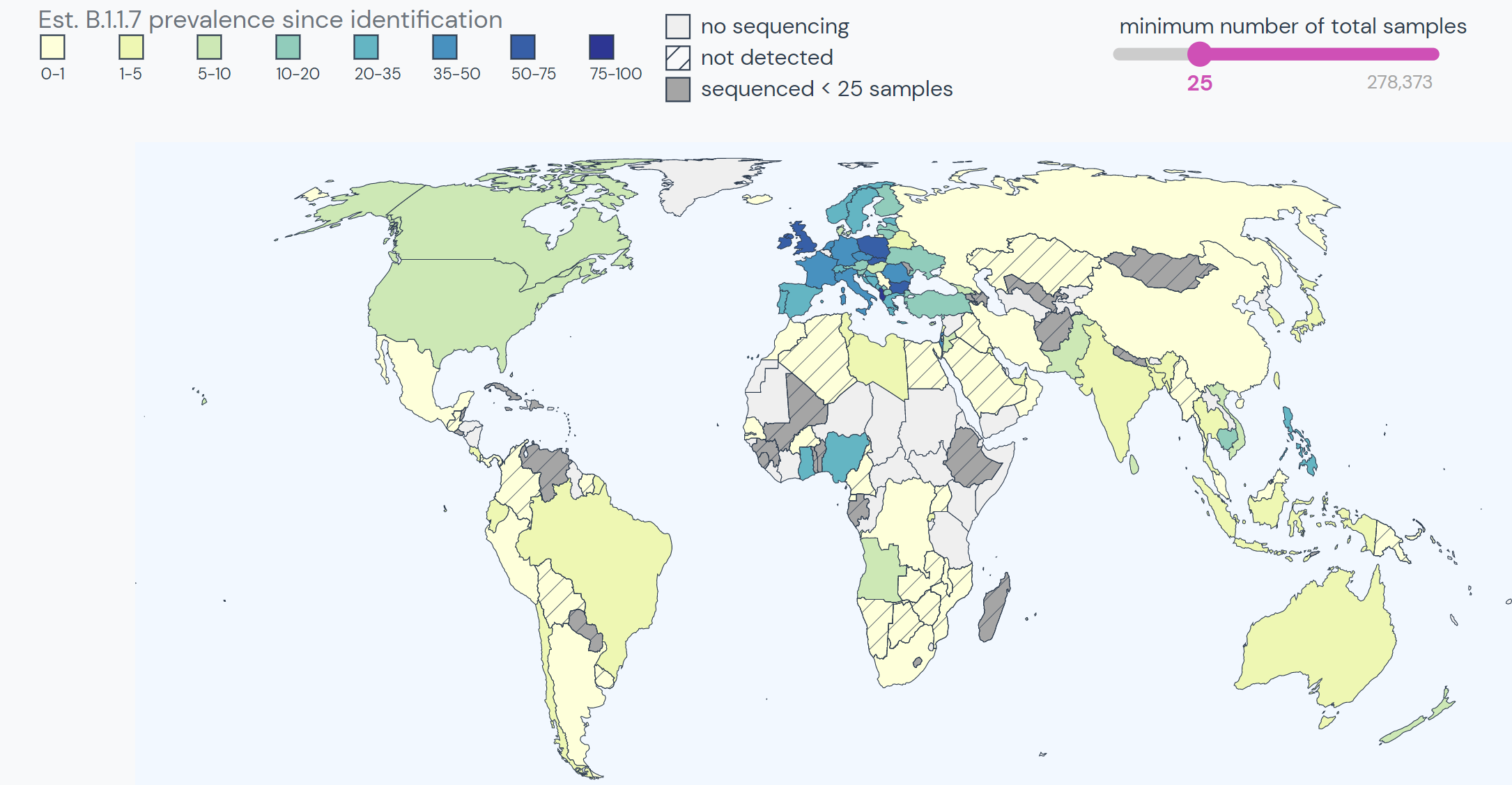
The variant has spread effectively through Europe. Wherever it is detected, B.1.1.7 tends to become the dominant strain very quickly. For example, B.1.1.7 made up a little over 5% of cases in Norway in early January. By mid-February, the variant accounted for 50% of sequenced samples. Today, the variant accounts for over 80% of sequenced samples.
The epidemic chart of worldwide average daily B.1.1.7 prevalence from the time of its detection is alarming for how closely it resembles a rocket launching:
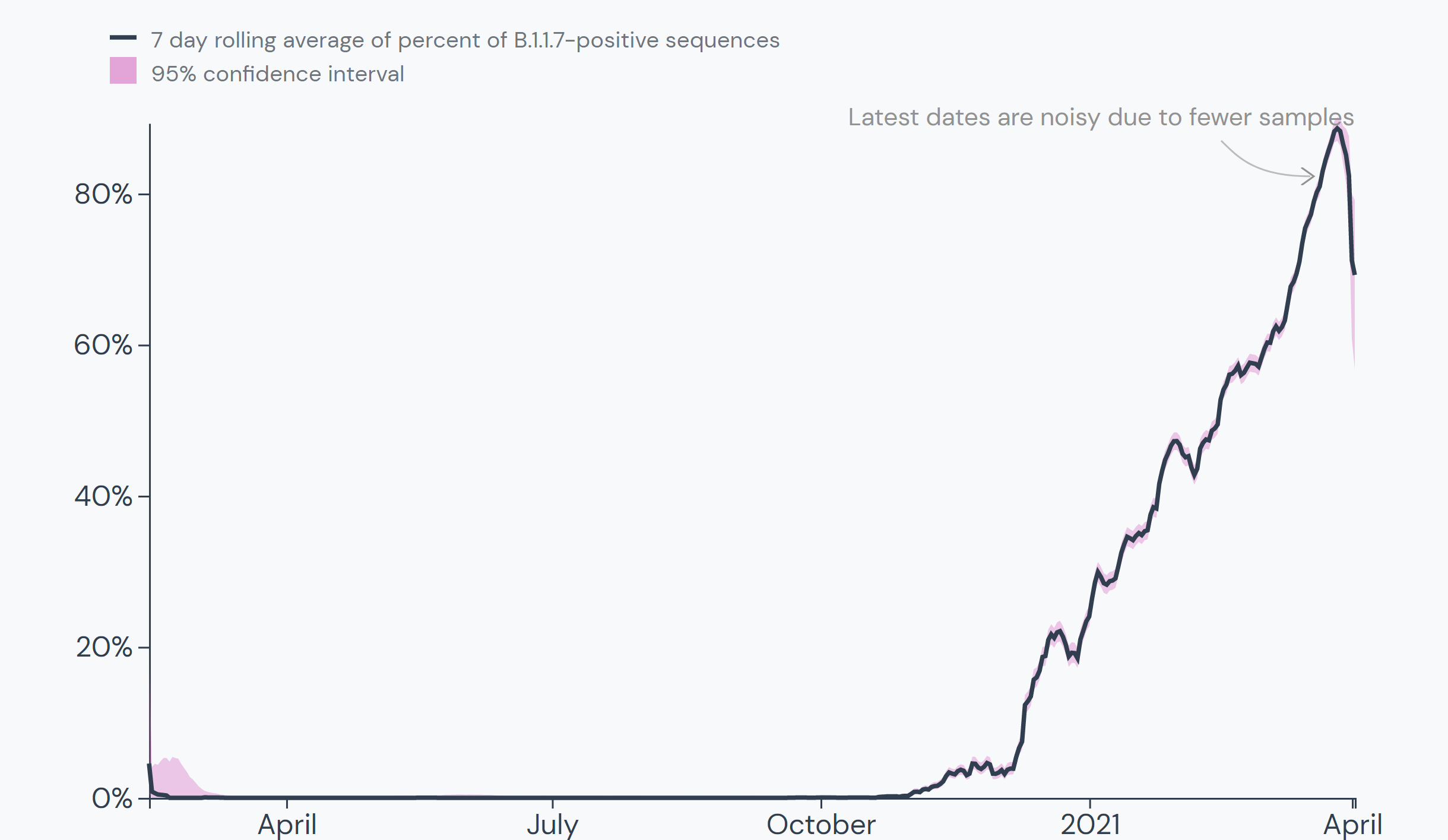
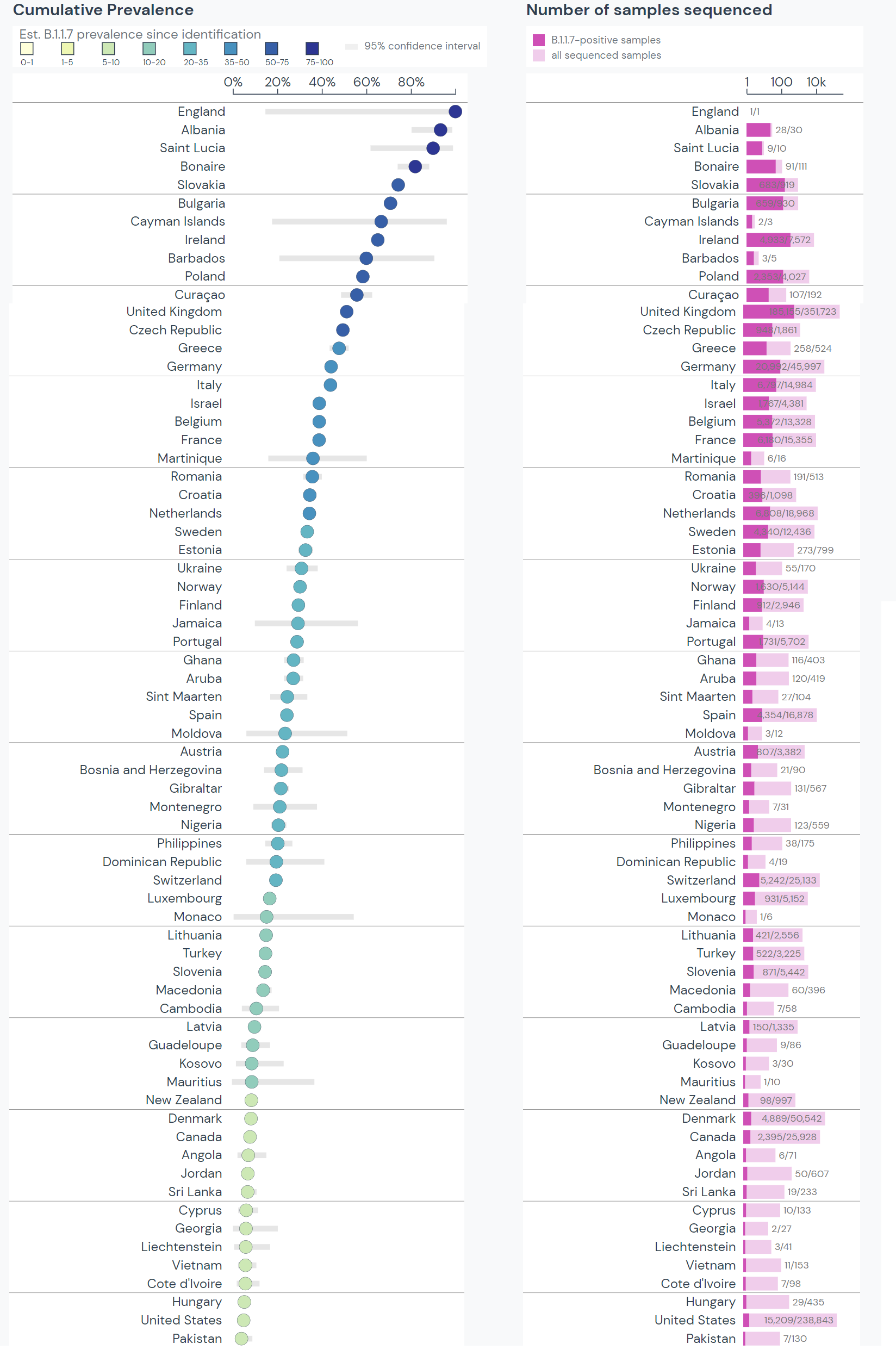
Around the world, 68 countries have experienced a cumulative B.1.1.7 prevalence over 5%. Estimates for many countries are unreliable, however, as they have sequenced few samples since first detection. Confidence intervals are presented for cumulative prevalence, indicated by the grey bars.
B.1.1.7 arrived in the US at the end of November 2020. Since then, the variant has spread through all US states; however, much of the East coast continues to experience the highest prevalence. Research has suggested a transmission pattern between Florida, Georgia, Minnesota, Michigan, and North Carolina.
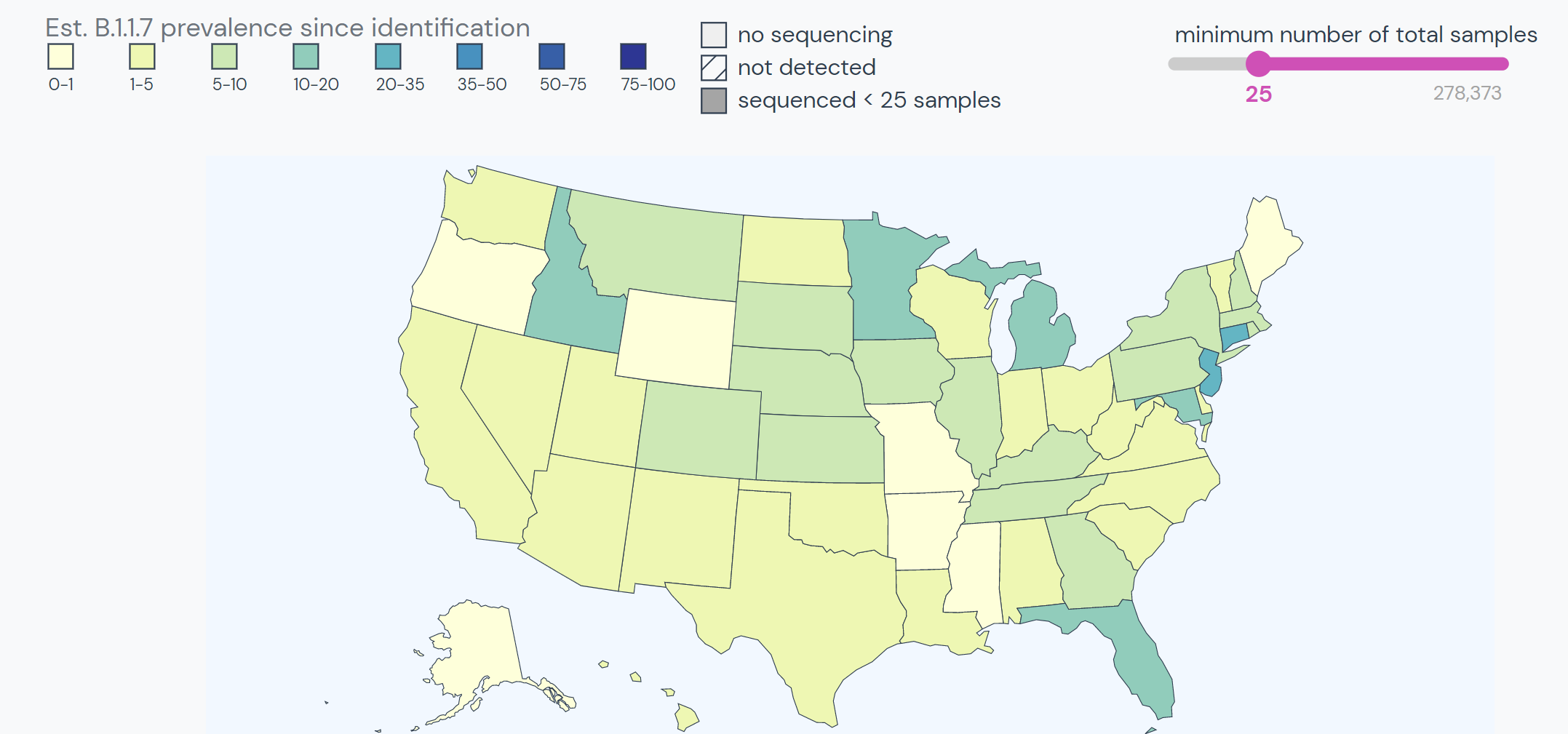
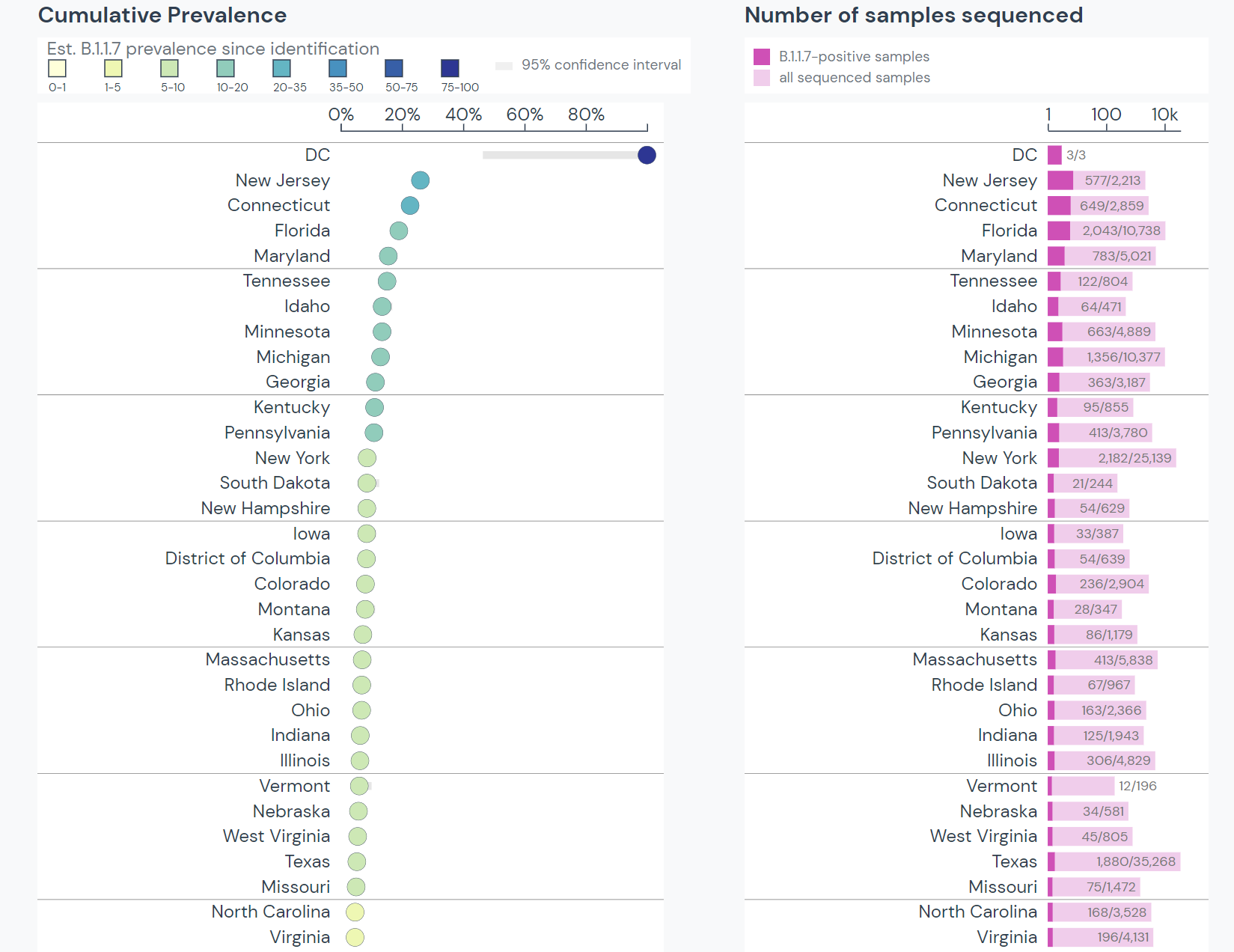
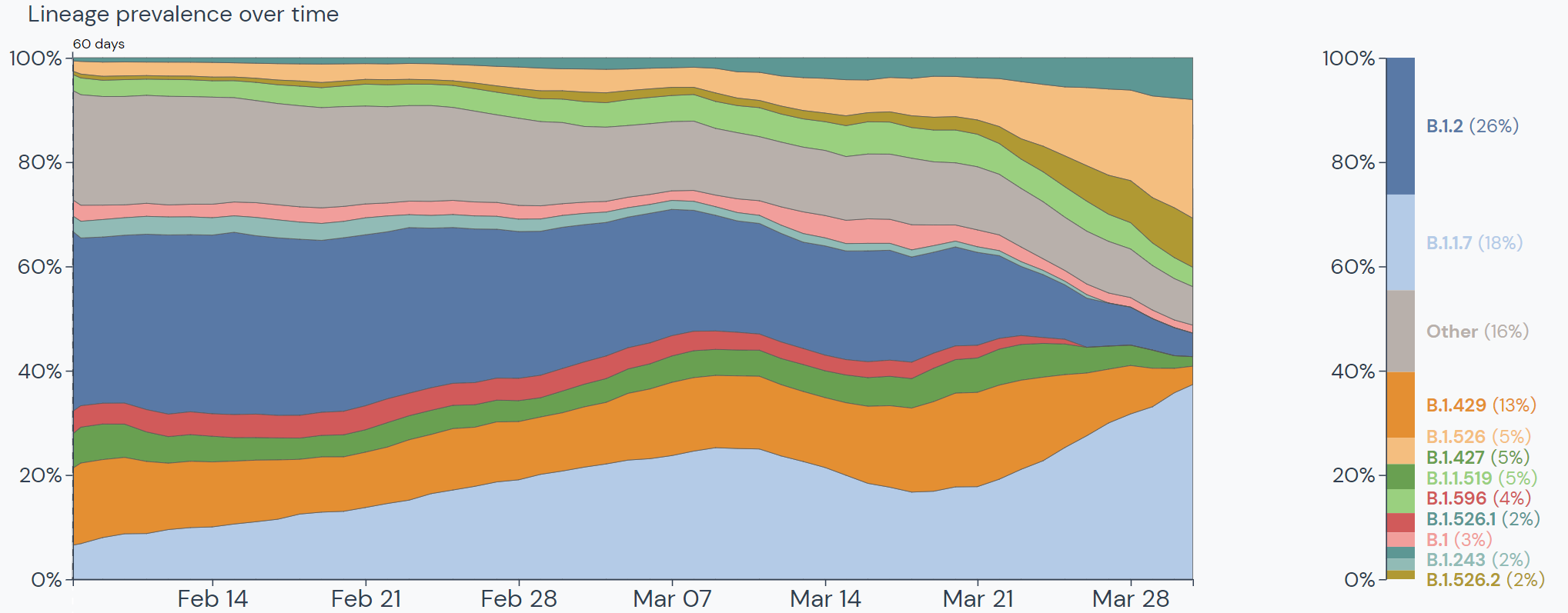
Between mid-January and mid-March, the prevalence of B.1.1.7 grew from 76 cases in 12 states to over 7,500 cases in all 50 states. B.1.1.7 makes up 18% of lineages sequenced in the US. The only lineage currently outcompeting B.1.1.7 in the US is B.1.2, which is the most prevalent lineage in the US but does not have the characteristics of a Variant of Interest/Concern.
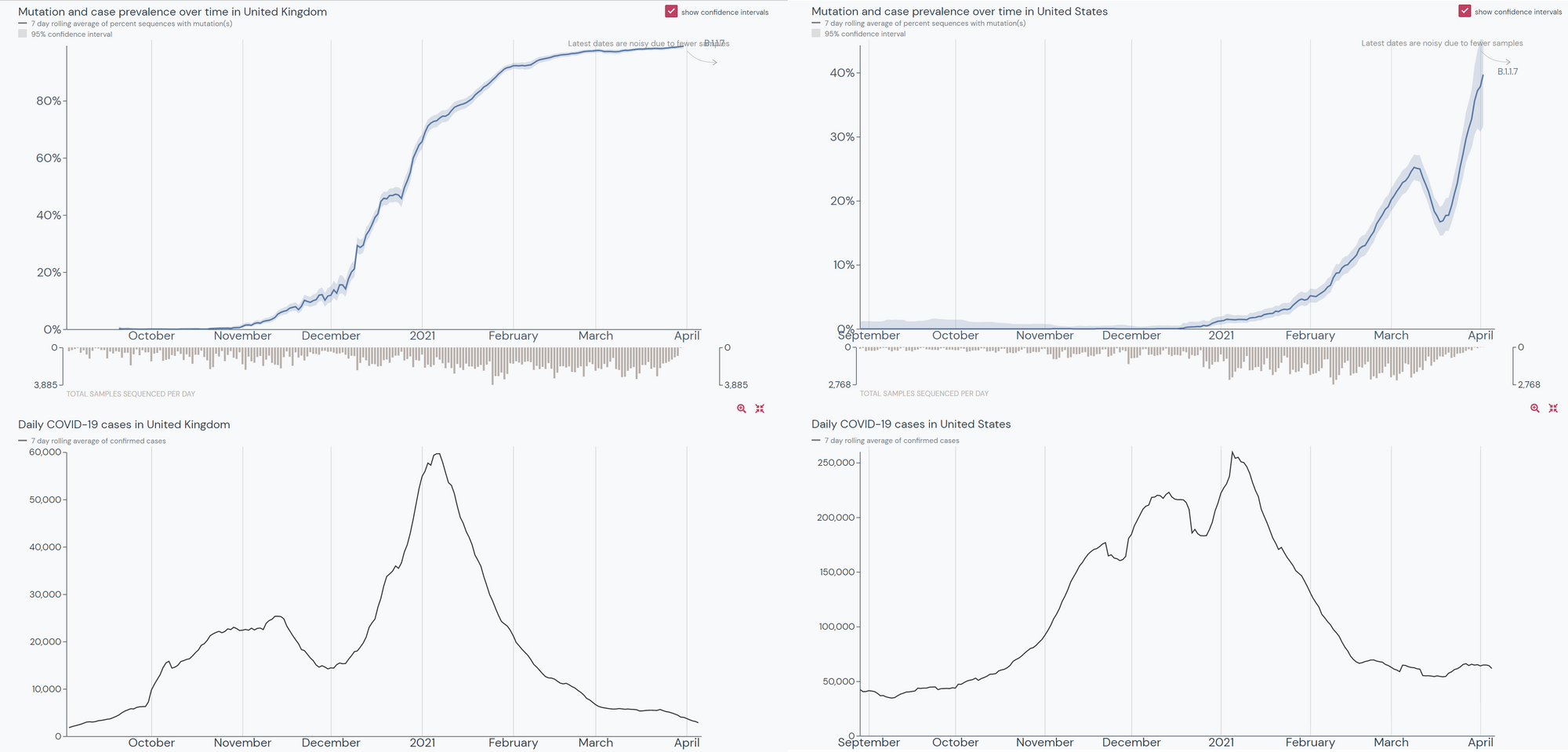
Sharply rising prevalence of the B.1.1.7 lineage in the UK coincides with a spike in overall COVID-19 cases in the country. This is not as evident in the US where B.1.1.7 has not been as prevalent for as long.
B.1.1.7 has been forecasted to become the dominant SARS-CoV-2 lineage in the United States, meaning cumulative prevalence will likely rise over 50%. The impact of vaccination as well as other newly emerging lineages remains unclear. The only thing that is certain is that the snapshot is constantly changing. As the more contagious coronavirus variant spreads throughout the world, we can continue to track global and national prevalence rates, epidemiological patterns, and even mutation patterns within genomes using Outbreak.info. Continued surveillance of the variant, where it is spreading, and its effect on the trajectory of COVID-19 cases for regions around the world is key to fighting the pandemic.
